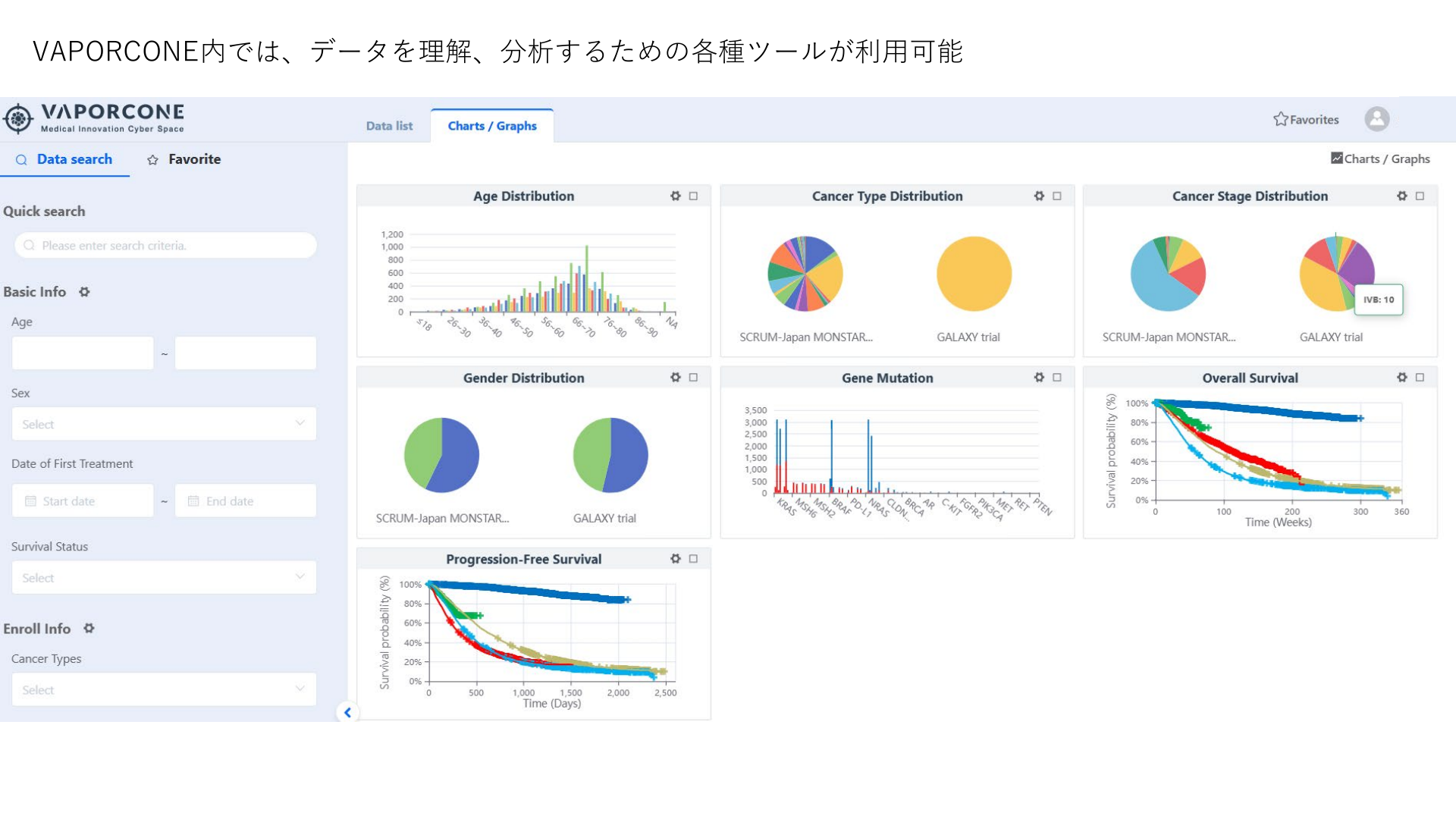
Clinical feasibility, treatment landscape, and biomarker availability
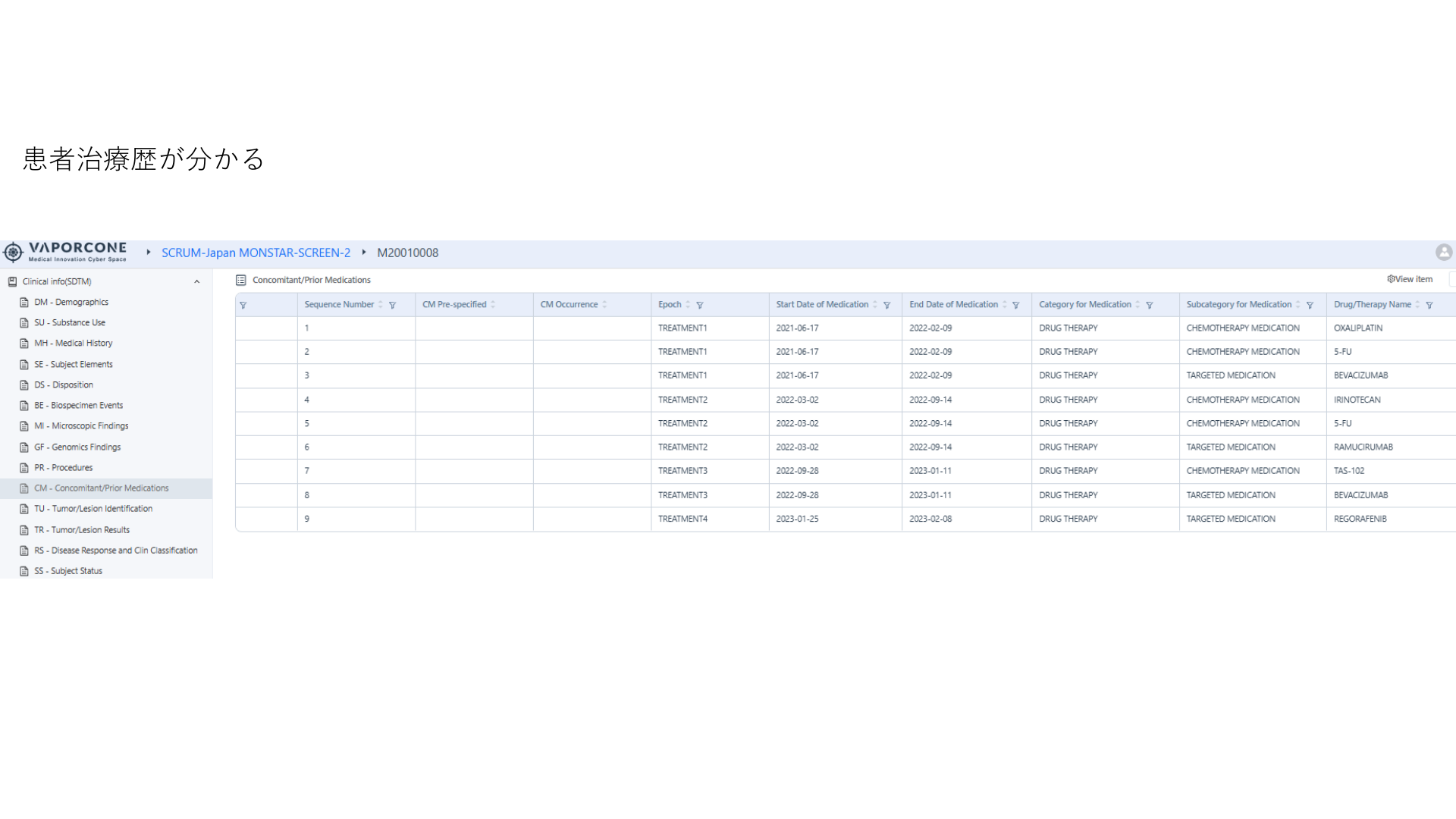
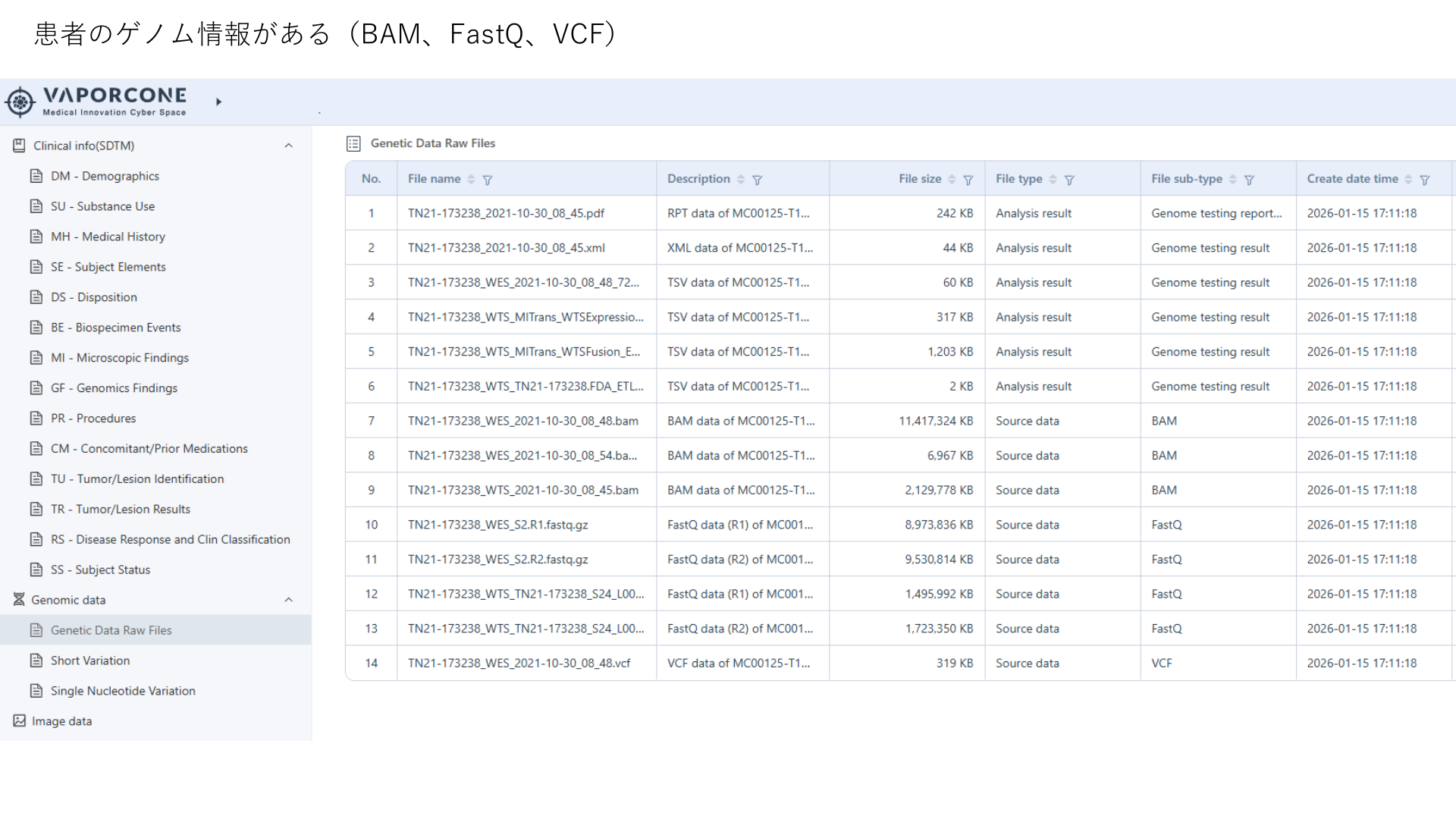
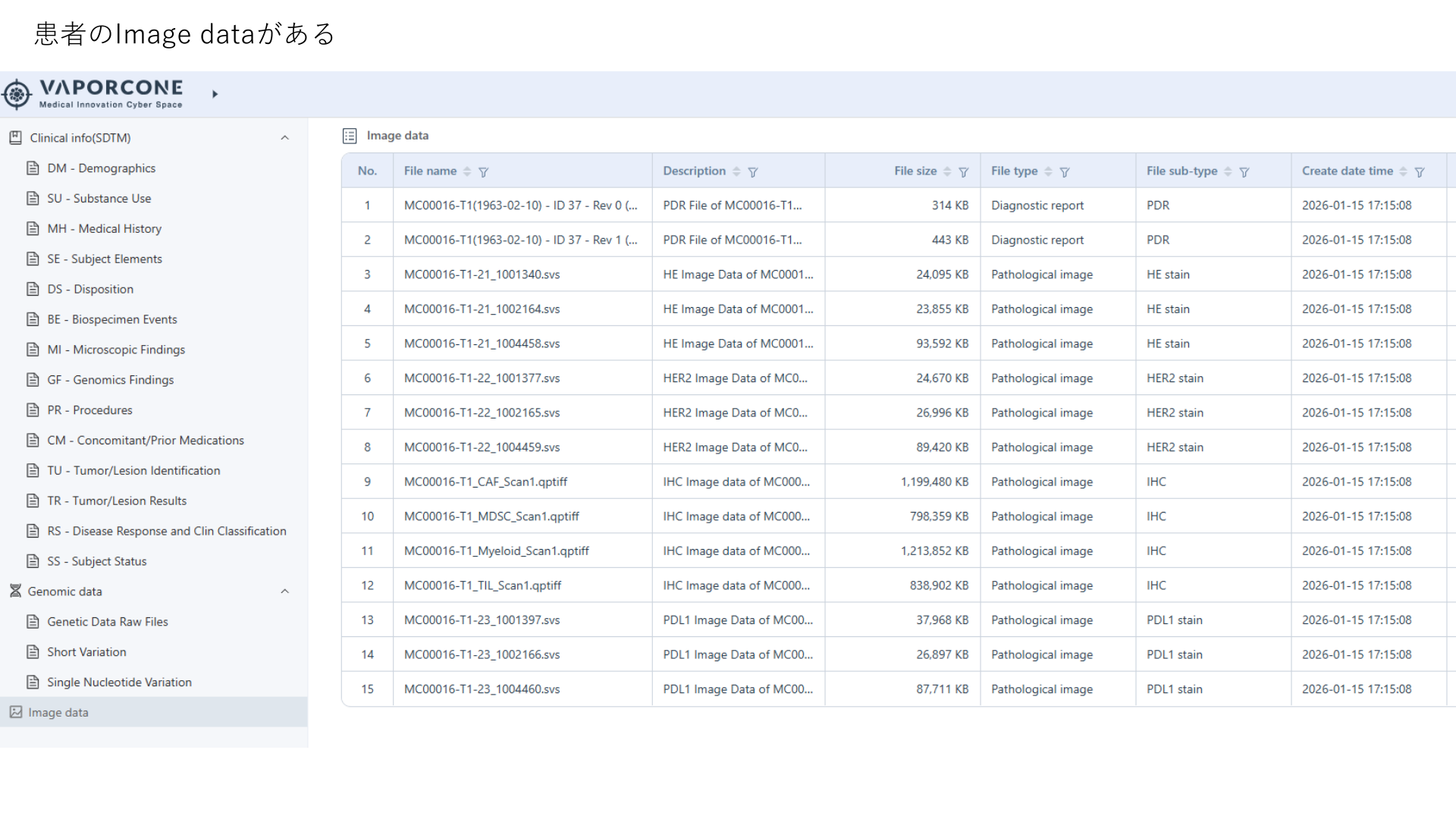
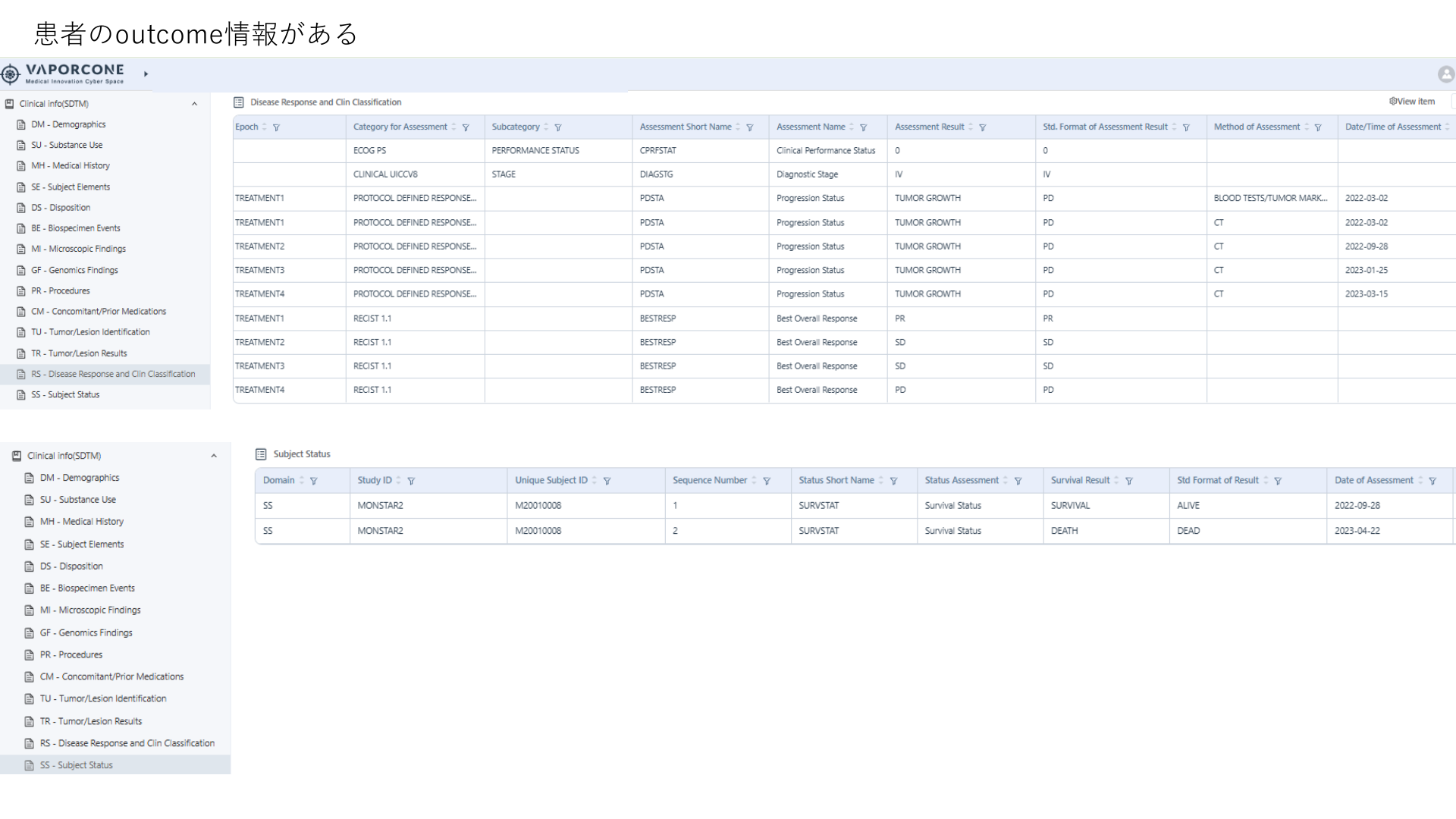
Cyber Oncology answers questions that start from routine clinical care: tumor type, stage, biomarker testing, line of therapy, medication exposure, labs, response, and safety.
Indication & Market Understanding
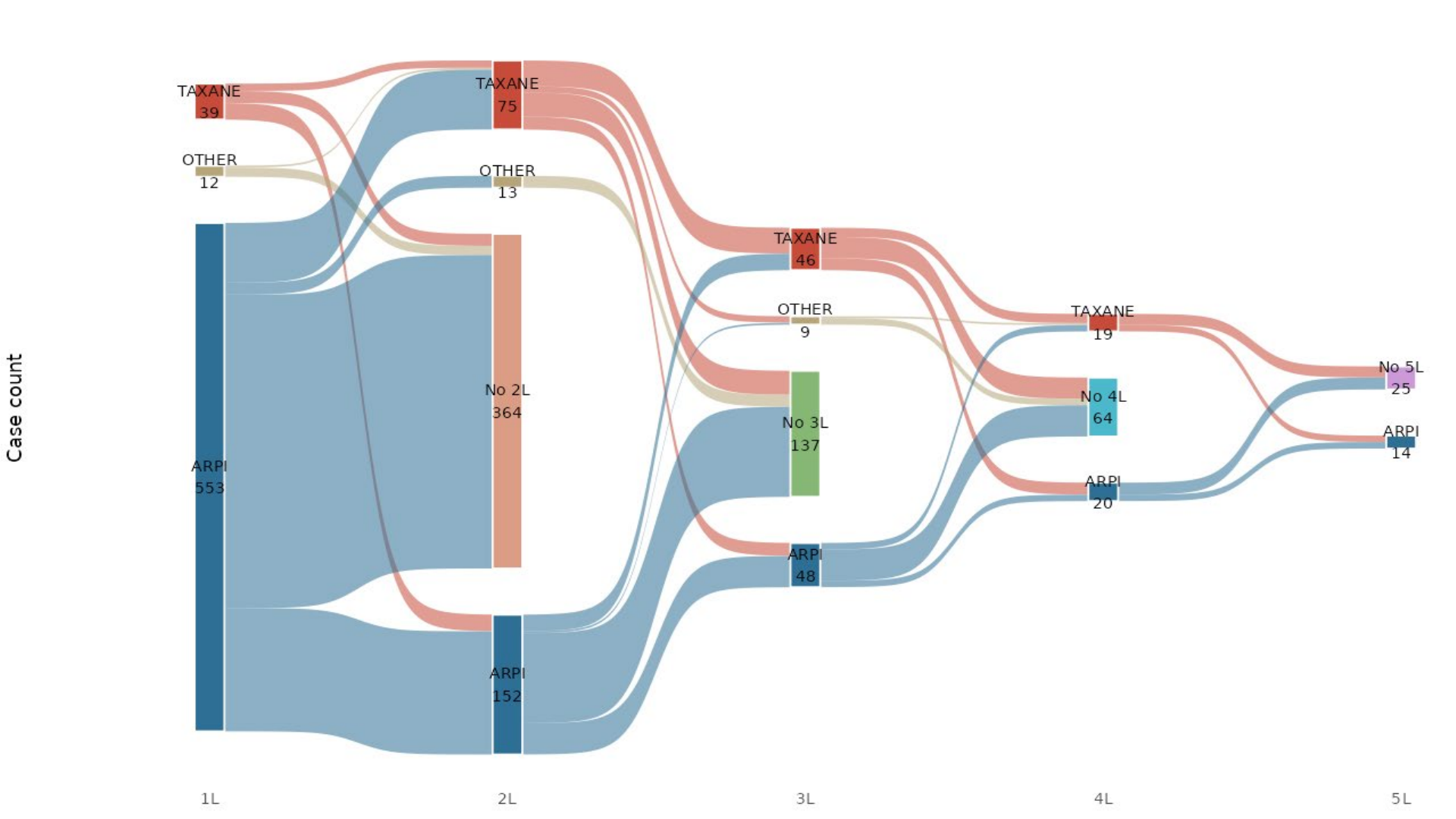
Treatment landscape, line-of-therapy sequencing, biomarker testing rates, addressable populations, and Japan-specific standard of care.
Trial Design & Feasibility
Patient counts against inclusion/exclusion criteria, endpoint availability, and site concentration for feasible enrollment planning.
Post-Approval & Label Expansion
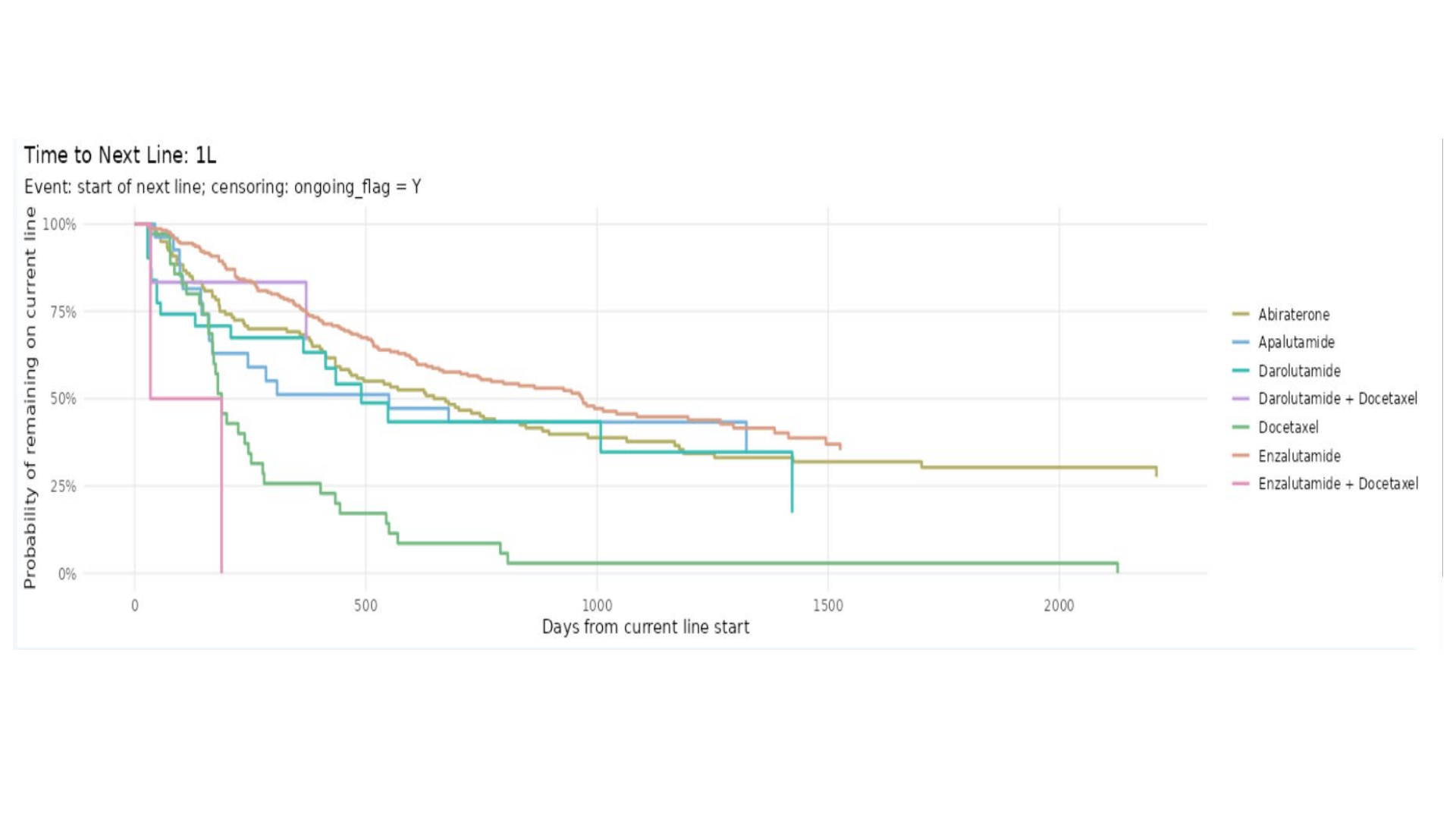
Real-world outcomes by therapy line, CTCAE-graded safety, HEOR endpoints, and subpopulation evidence.
Biomarker test results by tumor type
Approximate screened patients per biomarker, sourced from NTT catalog (Dec. 2025). This table was moved from Data Catalog to keep analysis examples together.
| Biomarker | BC | NSCLC | CRC | PDAC | GC | HCC | EC | OC | SCLC | BTC |
|---|---|---|---|---|---|---|---|---|---|---|
| ER | 3,880 | - | - | - | - | - | - | 260 | - | - |
| PR | 3,760 | - | - | - | - | - | - | 140 | - | - |
| HER2 | 5,480 | - | <100 | - | 800 | - | - | - | - | - |
| Ki-67 | 2,160 | <100 | - | - | - | - | - | - | <100 | - |
| EGFR | <100 | 1,360 | <100 | - | - | - | - | - | <100 | - |
| ALK fusion | - | 1,060 | - | - | - | - | - | - | <100 | - |
| PD-L1 | <100 | 1,260 | - | - | <100 | - | <100 | <100 | <100 | - |
| ROS1 fusion | - | 750 | - | - | - | - | - | - | - | - |
| BRAF | - | 610 | 440 | - | - | - | - | - | - | - |
| KRAS | - | <100 | 500 | - | - | - | - | - | - | - |
| MSI | <100 | - | 160 | <100 | 110 | <100 | <100 | <100 | - | <100 |
| MIB-1 | 1,670 | - | <100 | - | <100 | - | <100 | <100 | <100 | - |
Biomarkers have been obtained for approximately half of the cases in the patient totals. Expansion is ongoing.
NSCLC - Osimertinib 1L (EGFR+)
11 sites - 2018-2022 - Mar. 2026 estimate
| Cohort filter | Patients |
|---|---|
| Total NSCLC | 4,497 |
| Advanced/metastatic (IIIb-IV) | 2,719 |
| Tested for EGFR | 1,391 |
| EGFR(+) | 407 |
| Common EGFR | 281 |
| EGFR TKI received | 279 |
| Osimertinib as 1L | 246 |
| Osimertinib 1L monotherapy after Apr. 2018 | 246 |
BC - CDK4/6 inhibitor (HR+/HER2-)
11 sites - 2018-2023 - Mar. 2026 estimate
| Cohort filter | Patients |
|---|---|
| Total BC | 10,247 |
| Advanced/metastatic (IIIb-IV) | 1,247 |
| Tested for HR | 736 |
| HR(+) | 558 |
| Tested for HER2 | 880 |
| HER2(-) | 672 |
| HR(+) & HER2(-) | 413 |
| CDK4/6 inhibitor after HR/HER2 test | 222 |
Breast cancer - stage x biomarker testing matrix
Mar. 2026 estimate only. Positive biomarker counts require follow-up discussion because additional collection incurs cost.
| Stage | Patients | ER tested | PR tested | HER2 tested | Ki-67 tested |
|---|---|---|---|---|---|
| Stage I | 4,776 | 2,887 | 2,815 | 3,484 | 1,738 |
| Stage II | 3,860 | 2,312 | 2,290 | 2,806 | 1,379 |
| Stage III | 944 | 559 | 548 | 702 | 389 |
| Stage IV | 665 | 347 | 343 | 470 | 233 |
Prostate cancer - 15-facility projected matrix
15-facility projected matrix. Includes patients who underwent surgery or radiation therapy. Medication estimates are calculated from the 7-facility subset and projected to the 15-facility network.
| Stage | Patients | Anti-androgens | Taxane | BMA | Pluvicto | |||
|---|---|---|---|---|---|---|---|---|
| Any | Gen 1 | Gen 2 | CYP17 inh. | |||||
| ALL | 15,730 | 12,220 | 10,940 | 2,570 | 1,230 | 1,050 | 1,750 | 0 |
| Stage 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Stage I | 5,090 | 3,510 | 3,450 | 120 | 60 | 60 | 60 | 0 |
| Stage II | 3,280 | 2,340 | 2,340 | 120 | 60 | 60 | 60 | 0 |
| Stage III | 2,750 | 2,220 | 2,160 | 290 | 60 | 120 | 60 | 0 |
| Stage IV | 4,330 | 3,920 | 2,750 | 2,050 | 1,110 | 760 | 1,520 | 0 |
| Other | 350 | 290 | 290 | 60 | 0 | 0 | 0 | 0 |